Step 2: Check and Run the Data Cleaning Script
Cleaning the raw data can be done by simply running an R script (attached in the Toolkit). Here, I will explain each part of the script so you can revise it to fit your own survey.
See this guide on creating and executing an R file in R Studio.
Importing and Preparing Raw Data
raw <- read.csv('insert file path here')Import the raw data by using the read.csv() function.
raw <- replace(raw, raw == '', NA)
raw$ego_ID <- 10000 + 1:nrow(raw)Replace all empty values with NA by using the replace() function.
Egor requires numeric IDs for egos and alters. Here, I assigned numeric ego IDs (ego_ID) from 10001 to 10000 + (response number) to each response.
Creating an Index for Name Generator Questions
# Creating an index for identifying questions and ChoiceTextEntry values.
qid_varnames = c("Q1_NG_IM1", "Q2_NG_AC1") # Insert variable names of the name generators.
qid_varqids = c("QID1", "QID2") # Insert QIDs of the name generators.
a <- 10 # Insert the number of fields available in each name generator.
b <- 2 # Insert the number of name generator questions.
c <- a * bHere, we are going to create an index for identifying questions and ChoiceTextEntry values in the raw data.
- Insert variable names (available in the first row of the raw data) in the
qid_varnameslist. - Insert variable QIDs (available in the third row when saving the data in a CSV form) in the
qid_varqidslist. - Assign the number of fields available in each name generator as the value
a. In the sample survey, each IM and AC generator had 10 available fields, so I assigned 10 here. - Assign the number of name generator questions as the value
b. In the sample survey, there were two name generator questions (IM and AC generators), so the value ofbis 2.
qid <- data.frame(matrix(ncol = 3, nrow = c))
colnames(qid) <- c("QID1", "QID2", "QID3")
counter <- 1
for (m in 1:b) {
for (n in 1:a) {
qid$QID1[counter] <- paste0(qid_varnames[m], "_", n)
qid$QID2[counter] <- counter
qid$QID3[counter] <- paste0("{q://", qid_varqids[m], "/ChoiceTextEntryValue/", n, "}")
counter <- counter + 1
}
}You do not have to change anything from the above part. This part of the script will create a data frame that will be used as an index for coding the raw data. The data frame created from this script will look like:
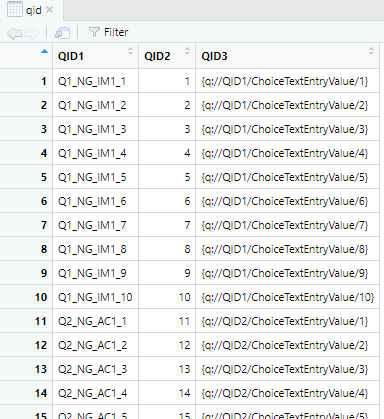
The QID index data frame produced by the script
Creating an Index for Density Questions
Next, we will create another data frame, which will be used as an index for coding the density (or alter-alter tie) data.
# Creating an index for identifying density questions and numbers
k <- nrow(qid)
h <- which(colnames(raw) == "Q9_Density_IM1_1") # Insert the first density question's variable name here.Insert the variable name of the first density question (available in the first row of the raw data). In this sample survey, the first variable's name was Q9_Density_IM1_1.
data <- list()
for (i in 1:(k-1)) {
for (j in (i+1):k) {
data <- c(data, list(c(i, j)))
}
}
densityid <- do.call(rbind, data)
colnames(densityid) <- c('fromid', 'toid')
densityid <- generate_densityid(k)
densityid <- as.data.frame(densityid)
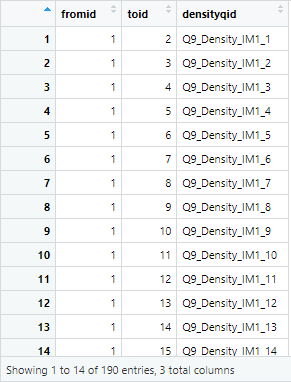
densityid$densityqid <- colnames(raw)[h:(h - 1 + ((k * (k-1)) / 2))]You don't have to change any part of this script. This script will generate a data frame (an index for coding the density or alter-alter tie data) that looks like this:

The densityid index data frame produced by the script
Creating the Alters Dataset
Now, we are going to make data frames for creating an egor object (threefiles_to_egor). Three files are required to make an egor object: 1) ego data, 2) alters data, and 3) edge data (alter-alter tie or density data).
We are going to make alters data first.
# Create an empty data frame "alter." Revise the structure to fit your data.
alter_raw <- data.frame(
response_ID = character(),
ego_ID = numeric(),
alter_name = character(),
alter_ID_char = character(),
alter_ID = numeric(),
alter_relationship = character(),
alter_gender = character(),
alter_group = character(),
alter_sampled = character(),
stringsAsFactors = FALSE
)This script makes an empty data frame that will be used for storing alters data. Create columns that fit your alter information structure. In this sample survey, we had two name interpreters ("Relationship Type" and "Gender"). So there are alter_relationship and alter_gender columns included in this structure. If you have used more interpreters and need more columns to store data, include those here.
alter_group is necessary when using multiple name generators (e.g., "Important Matters" and "Academic & Career Matters"). This column will store information on which group ("Important Matters," or "Academic & Career Matters," or both) each listed alter is affiliated.
Other parts (response_ID, ego_ID, alter_name, alter_ID_char, alter_ID, alter_sampled, stringsAsFactors) should be left as they are now, since these are for storing the basic information of alters and egos.
# Create an empty data frame "alter_temp." Revise the structure to fit your data.
num <- nrow(raw)
alter_temp <- data.frame(
response_ID = character(num),
ego_ID = numeric(num),
alter_name = character(num),
alter_ID_char = character(num),
alter_ID = numeric(num),
alter_relationship = character(num),
alter_gender = character(num),
alter_group = character(num),
alter_sampled = character(num),
stringsAsFactors = FALSE
)This script makes an empty data frame that will be used for storing temporary alters data while cleaning. Make this data frame structure look the same as alter_raw. If you added more columns in alter_raw, the same columns should be added in this data frame as well.
The below script is for cleaning the data: It stores the required information for alters data.
# Identifying variables & recording the responses.
# Note: Adjust the column/variable names (e.g., raw$SN_showsample_DO) to align with your dataset.
for (i in seq_len(nrow(qid))) {
qid1_val <- qid[i, "QID1"]
qid2_val <- qid[i, "QID2"]
qid3_val <- qid[i, "QID3"]
if (!is.null(raw[[qid1_val]])) {
alter_temp$response_ID <- raw$ResponseId # Stores Qualtrics response IDs
alter_temp$ego_ID <- raw$ego_ID # Stores ego IDs generated in the previous step
alter_temp$alter_name <- raw[[qid1_val]] # Stores names listed in the name generators
alter_temp$alter_ID_char <- paste(raw$ResponseId, "_alter", qid2_val, sep = "")
# Assigns alter IDs in character (responseID_alter_number)
alter_temp$alter_ID <- paste((raw$ego_ID) * 100 + qid2_val, sep = "")
# Assigns alter IDs in numeric values ("egoID"*100 + number)You don't have to change anything in the above part. This script is for saving basic information (response ID of each respondent ('ego'), alter names and IDs).
alter_temp$alter_relationship <- raw[[paste("SN_relationship_", qid2_val, sep = "")]]
# Stores information provided in the "SN_relationship" column.
alter_temp$alter_gender <- raw[[paste("SN_gender_", qid2_val, sep = "")]]
# Stores information provided in the "SN_gender" column.Revise this part of the script to fit your data. In the sample survey, there were two name interpreters, so there are two lines of script. Each script stores data in the SN_relationship column and SN_gender column in the raw data.
alter_temp$alter_group <- ifelse(
!is.na(raw$Q3_NG_IM_AC_select) &
raw$Q3_NG_IM_AC_select != "" &
grepl(paste0("\\$\\", qid3_val), raw$Q3_NG_IM_AC_select),
"IMAC",
ifelse(qid2_val > (a * (b - 1)), "AC", "IM")
)
# Stores information about alter's group affiliation
# (e.g., 'important matters', 'academic and career matters', or both)This line stores alter's group information (e.g., whether the alter was listed in the 'important matters' list, 'academic and career matters' list, or both lists). This information could be found in the overlap measure we created in the survey (see Step 3). The variable name for this question was Q3_NG_IM_AC_select, so I inserted the variable name in the underline spots. Revise the variable name to fit your data.
alter_temp$alter_sampled <- ifelse(
!is.na(raw$Q6_Random_DO) &
raw$Q6_Random_DO != "" &
grepl(paste0("\\$\\", qid3_val), raw$Q6_Random_DO),
"Yes",
"No"
)
# Stores information about whether the alter is sampled for the name interpreters.This line stores information about whether the alter is sampled for the name interpreters. This information could be found in the [sample list variable name]_DO column in the raw data. Because I named the sample list question as Q6_Random, the column name is Q6_Random_DO. (DO is short for Display Order.) Revise the variable name to fit your data.
alter_raw <- rbind(alter_raw, alter_temp)
}
}
alter_raw <- replace(alter_raw, alter_raw == '', NA)
alter_clean <- alter_raw[complete.cases(alter_raw$alter_name), ]You don't have to change anything in the above part of the script.
The above script will generate "alters data" as shown below:

The alter_clean data frame produced by the cleaning script
Creating the Density (Edge / Alter-Alter Tie) Dataset
Next, we are going to create a density ("edge" or "alter-alter tie") dataset. This part of the script is fully automated, and you won't have to change anything from this script.
# Creating density (edge or alter-alter tie) dataset
alter_ties_raw <- data.frame(
response_ID = character(),
ego_ID = numeric(),
from = character(),
to = character(),
weight = numeric(),
stringsAsFactors = FALSE
)
alter_ties_temp <- data.frame(
response_ID = character(num),
ego_ID = numeric(num),
from = character(num),
to = character(num),
weight = numeric(num),
stringsAsFactors = FALSE
)
for (j in seq_len(nrow(densityid))) {
denid_val1 <- densityid[j, "densityqid"]
denid_val2 <- densityid[j, "fromid"]
denid_val3 <- densityid[j, "toid"]
if (!is.null(raw[[denid_val1]])) {
alter_ties_temp$response_ID <- raw$ResponseId
alter_ties_temp$ego_ID <- raw$ego_ID
alter_ties_temp$from <- ifelse(
!is.na(raw[[denid_val1]]),
paste((raw$ego_ID * 100) + denid_val2, sep = ""),
NA
)
alter_ties_temp$to <- ifelse(
!is.na(raw[[denid_val1]]),
paste((raw$ego_ID * 100) + denid_val3, sep = ""),
NA
)
alter_ties_temp$weight <- ifelse(!is.na(raw[[denid_val1]]), 1, NA)
alter_ties_raw <- rbind(alter_ties_raw, alter_ties_temp)
}
}
alter_ties_raw <- replace(alter_ties_raw, alter_ties_raw == '', NA)
alter_ties_clean <- alter_ties_raw[complete.cases(alter_ties_raw$weight), ]The above script will generate the alter-alter ties data frame as shown below:
The alter_ties_clean data frame produced by the cleaning script
response_ID and ego_ID are the IDs of the respondent ('ego'). The from and to columns have alter IDs, representing pairs of alters who know/hang out/talk to each other. weight is 1 in this dataset because we did not ask questions about closeness or tie strength between listed alters.
Saving the Created Datasets
# Saving the created datasets
ego <- raw[, c("ego_ID", "ResponseId")]
colnames(ego) <- c("ego_ID", "response_ID")The above script creates ego data.
write.csv(ego, "ego_data.csv")
write.csv(alter_clean, "alter_attr.csv")
write.csv(alter_ties_clean, "alter_ties.csv")Finally, this script stores 1) ego data, 2) alters data, and 3) edge data (alter-alter tie or density data) as CSV files. If you want to learn how to create an egor object, click here.